Nsp1 facilitates SARS-CoV-2 replication through calcineurin-NFAT signaling
Wai-Yin Lui, Chon Phin Ong, Pak-Hin Hinson Cheung, Zi-Wei Ye, Chi-Ping Chan, Kelvin Kai-Wang To, Kit-San Yuen, Dong-Yan Jin
mBio, doi:10.1128/mbio.00392-24
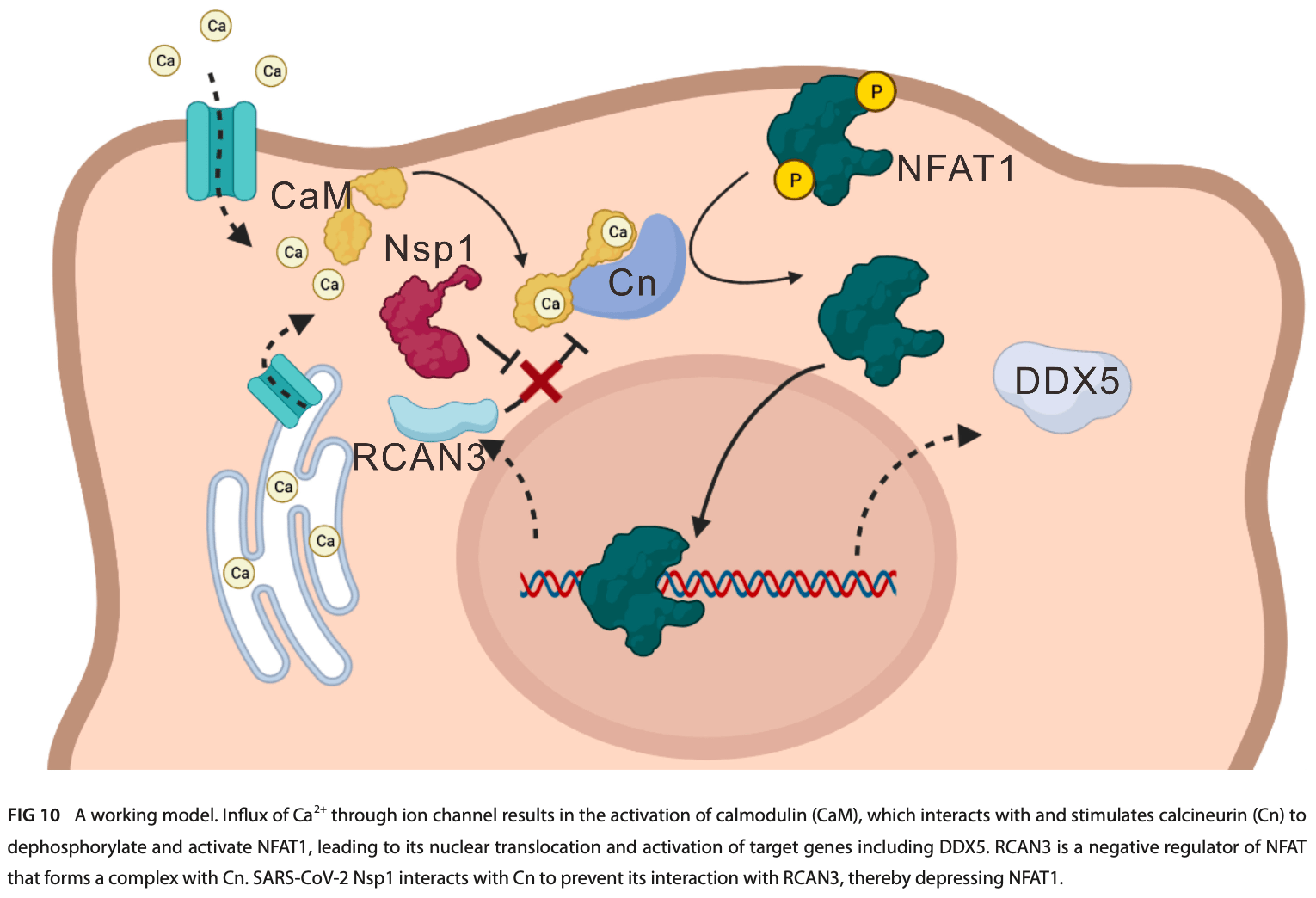
SARS-CoV-2, the causative agent of COVID-19, has been intensely studied in search of effective antiviral treatments. The immunosuppressant cyclosporine A (CsA) has been suggested to be a pan-coronavirus inhibitor, yet its underlying mechanism remained largely unknown. Here, we found that non-structural protein 1 (Nsp1) of SARS-CoV-2 usurped CsA-suppressed nuclear factor of activated T cells (NFAT) signal ing to drive the expression of cellular DEAD-box helicase 5 (DDX5), which facilitates viral replication. Nsp1 interacted with calcineurin A (CnA) to displace the regulatory protein regulator of calcineurin 3 (RCAN3) of CnA for NFAT activation. The influence of NFAT activation on SARS-CoV-2 replication was also validated by using the Nsp1-deficient mutant virus. Calcineurin inhibitors, such as CsA and VIVIT, inhibited SARS-CoV-2 replication and exhibited synergistic antiviral effects when used in combination with nirmatrelvir. Our study delineated the molecular mechanism of CsA-mediated inhibition of SARS-CoV-2 replication and the anti-SARS-CoV-2 action of calcineurin inhibitors. IMPORTANCE Cyclosporine A (CsA), commonly used to inhibit immune responses, is also known to have anti-SARS-CoV-2 activity, but its mode of action remains elusive. Here, we provide a model to explain how CsA antagonizes SARS-CoV-2 through three critical proteins: DDX5, NFAT1, and Nsp1. DDX5 is a cellular facilitator of SARS-CoV-2 replication, and NFAT1 controls the production of DDX5. Nsp1 is a viral protein absent from the mature viral particle and capable of activating the function of NFAT1 and DDX5. CsA and similar agents suppress Nsp1, NFAT1, and DDX5 to exert their anti-SARS-CoV-2 activity either alone or in combination with Paxlovid.
DIRECT CONTRIBUTION This article is a direct contribution from Dong-Yan Jin, a Fellow of the American Academy of Microbiology, who arranged for and secured reviews by Haitao Guo, University of Pittsburgh, and Jianming Qiu, The University of Kansas Medical Center.
DATA AVAILABILITY All data reported in this paper will be shared by lead upon request.
References
Abaeva, Arhab, Miścicka, Hellen, Pestova, In vitro reconstitution of SARS-CoV-2 Nsp1-induced mRNA cleavage reveals the key roles of the N-terminal domain of Nsp1 and the RRM domain of eIF3g, Genes Dev,
doi:10.1101/gad.350829.123Afsar, Narayan, Akhtar, Das, Rahil et al., Drug targeting Nsp1-ribosomal complex shows antiviral activity against SARS-CoV-2, Elife,
doi:10.7554/eLife.74877Basic-Jukic, Juric, Furic-Cunko, Katalinic, Radic et al., Follow-up of renal transplant recipients after acute COVID-19-A prospective cohort single-center study, Immun Inflamm Dis,
doi:10.1038/s41598-022-14194-xBrewin, Mancao, Straathof, Karlsson, Samarasinghe et al., Generation of EBV-specific cytotoxic T cells that are resistant to calcineurin inhibitors for the treatment of posttransplanta tion lymphoproliferative disease, Blood,
doi:10.1182/blood-2009-07-228387Chan, Ip, Chu, Yip, Lo et al., Identification of Nsp1 gene as the target of SARS-CoV-2 real-time RT-PCR using nanopore whole-genome sequencing, J Med Virol,
doi:10.1002/jmv.26140Chan, Yuan, Kok, To, Chu et al., A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster, Lancet,
doi:10.1016/S0140-6736(20)30154-9Cheng, Ip, Chu, Tam, Chan et al., Rapid spread of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Omicron subvariant BA.2 in a single-source community outbreak, Clin Infect Dis,
doi:10.1093/cid/ciac203Cheung, Ye, Lui, Ong, Chan et al., Production of single-cycle infectious SARS-CoV-2 through a trans-complemented replicon, J Med Virol,
doi:10.1002/jmv.28057Dittmar, Lee, Whig, Segrist, Li et al., Drug repurposing screens reveal cell-type-specific entry pathways and FDA-approved drugs active against SARS-CoV-2, Cell Rep,
doi:10.1016/j.celrep.2021.108959Eckstein, Van Quill, Bui, Uusitalo, Brien, Cyclosporin a inhibits calcineurin/nuclear factor of activated T-cells signaling and induces apoptosis in retinoblastoma cells, Invest Ophthalmol Vis Sci,
doi:10.1016/j.cmi.2023.01.002Fenizia, Galbiati, Vanetti, Vago, Clerici et al., Cyclosporine A inhibits viral infection and release as well as cytokine production in lung cells by three SARS-CoV-2 variants, Microbiol Spectr,
doi:10.1128/spectrum.01504-21Fishbane, Hirsch, Nair, Special considerations for Paxlovid treatment among transplant recipients with SARS-CoV-2 infection, Am J Kidney Dis,
doi:10.1053/j.ajkd.2022.01.001Frolov, Agback, Palchevska, Dominguez, Lomzov et al., All domains of SARS-CoV-2 Nsp1 determine transla tional shutoff and cytotoxicity of the protein, J Virol,
doi:10.1128/jvi.01865-22Gonzalez-García, Rodríguez-García, Arranz, Ambrosioni, Bodro et al., Drug-drug interactions of ritonavir-boosted SARS-CoV-2 protease inhibitors in solid organ transplant recipients: experience from the initial use of lopinavir-ritonavir, Clin Microbiol Infect,
doi:10.1016/j.cmi.2023.01.002Guisado-Vasco, Valderas-Ortega, Mm, Santacruz, González-Cortijo et al., 166.201. receiving antiviral, antimalarials, glucocorticoids, or immunomodulation with tocilizumab cyclosporine: retrospective observational study (COQUIMA cohort), Research Article mBio Month,
doi:10.1016/j.eclinm.2020.100591Harari, Tahor, Rutsinsky, Meijer, Miller et al., Drivers of adaptive evolution during chronic SARS-CoV-2 infections, Nat Med,
doi:10.1038/s41591-022-01882-4Havkin-Solomon, Itzhaki, Joffe, Shaul, Dikstein, Selective translational control of cellular and viral mRNAs by RPS3 mRNA binding, Nucleic Acids Res,
doi:10.1093/nar/gkad269Kao, Orry, Palfreyman, Porton, Synergistic interactions of repurposed drugs that inhibit Nsp1, a major virulence factor for COVID-19, Sci Rep,
doi:10.1038/s41598-022-14194-xKumar, Ishida, Strilets, Cole, Lopez-Orozco et al., SARS-CoV-2 nonstructural protein 1 inhibits the interferon response by causing depletion of key host signaling factors, J Virol,
doi:10.1128/JVI.00266-21Lapointe, Grosely, Johnson, Wang, Fernández et al., Dynamic competition between SARS-CoV-2 NSP1 and mRNA on the human ribosome inhibits translation initiation, Proc Natl Acad Sci U S A,
doi:10.1073/pnas.2017715118Lin, Tang, Wei, Du, Chen et al., Genomic monitoring of SARS-CoV-2 uncovers an Nsp1 deletion variant that modulates type I interferon response, Cell Host Microbe,
doi:10.1016/j.chom.2021.01.015Liu, Stauft, Selvaraj, Chandrasekaran, 'agnillo et al., Intranasal delivery of a rationally attenuated SARS-CoV-2 is immuno genic and protective in Syrian hamsters, Nat Commun,
doi:10.1038/s41467-022-34571-4Lokugamage, Narayanan, Nakagawa, Terasaki, Ramirez et al., Middle East respiratory syndrome coronavirus Nsp1 inhibits host gene expression by selectively targeting mRNAs transcri bed in the nucleus while sparing mRNAs of cytoplasmic origin, J Virol,
doi:10.1128/JVI.01352-15Lui, Bharti, Wong, Jangra, Botelho et al., Suppression of cGAS-and RIG-I-mediated innate immune signaling by Epstein-Barr virus deubiquitinase BPLF1, PLoS Pathog,
doi:10.1371/journal.ppat.1011186Marques, Marques, Justino, Leukotrienes vs. montelu kast-activity, metabolism, and toxicity hints for repurposing, Pharma ceuticals,
doi:10.1038/s41592-022-01447-wMccallum, Walls, Sprouse, Bowen, Rosen et al., Molecular basis of immune evasion by the Delta and Kappa SARS-CoV-2 variants, Science,
doi:10.1126/science.abl8506Mendez, Ly, González-Sánchez, Hartenian, Ingolia et al., The N-terminal domain of SARS-CoV-2 Nsp1 plays key roles in suppression of cellular gene expression and preservation of viral gene expression, Cell Rep,
doi:10.1016/j.celrep.2021.109841Monticelli, Rao, Ong, Tang, Fan et al., Intranasal administration of a single dose of a candidate live attenuated vaccine derived from an NSP16-deficient SARS-CoV-2 strain confers sterilizing immunity in animals, Cell Mol Immunol,
doi:10.1038/s41423-022-00855-4Murgolo, Therien, Howell, Klein, Koeplinger et al., SARS-CoV-2 tropism, entry, replication, and propagation: considerations for drug discovery and development, PLoS Pathog,
doi:10.1016/j.cmi.2023.01.002Park, Yoo, Kim, The role of calcium-calcineurin-NFAT signaling pathway in health and autoimmune diseases, Front Immunol,
doi:10.1186/1478-811X-7-25Pfefferle, Schöpf, Kögl, Friedel, Müller et al., The SARScoronavirus-host Interactome: identification of cyclophilins as target for pan-coronavirus inhibitors, PLoS Pathog,
doi:10.1371/journal.ppat.1002331Poon, Constantinou, Lamoureux, Taylor, Topical cyclosporin A in the treatment of acute graft rejection: a randomized controlled trial, Clin Ex Ophthalmol,
doi:10.1111/j.1442-9071.2008.01784.xPrasad, Ahamad, Kanipakam, Gupta, Kumar, Simultane ous inhibition of SARS-CoV-2 entry pathways by cyclosporine, ACS Chem Neurosci,
doi:10.1021/acschemneuro.1c00019Sauerhering, Kuznetsova, Kupke, Meier, Halwe et al., Cyclosporin A reveals potent antiviral effects in preclinical models of SARS-CoV-2 infection, Am J Respir Crit Care Med,
doi:10.1164/rccm.202108-1830LESchubert, Karousis, Ban, Lapointe, Leibundgut et al., Universal features of Nsp1-mediated translational shutdown by coronaviruses, Mol Cell,
doi:10.1016/j.molcel.2023.09.002Schubert, Karousis, Jomaa, Scaiola, Echeverria et al., SARS-CoV-2 Nsp1 binds the ribosomal mRNA channel to inhibit translation, Nat Struct Mol Biol,
doi:10.1038/s41594-020-0511-8Sekhon, Shin, Kim, Jeong, Choi et al., Cyclophilin A-mediated mitigation of coronavirus SARS-CoV-2, Bioeng Transl Med,
doi:10.1002/btm2.10436Shin, Kim, Cho, Nguyen, Coupled feedback regulation of nuclear factor of activated T-cells (NFAT) modulates activation-induced cell death of T cells, Sci Rep,
doi:10.1038/s41598-019-46592-zSieber, Baumgrass, Novel inhibitors of the calcineurin/NFATc hub -alternatives to CsA and FK506?, Cell Commun Signal,
doi:10.1186/1478-811X-7-25Singh, Infectious complications in organ transplant recipients with the use of calcineurin-inhibitor agent-based immunosuppressive regimens, Curr Opin Infect Dis,
doi:10.1097/01.qco.0000172698.52408.beSlobodin, Sehrawat, Hayat, Zuckerman, Fraticelli et al., Capindependent translation and a precisely located RNA sequence enable SARS-CoV-2 to control host translation and escape anti-viral response, Nucleic Acids Res,
doi:10.1093/nar/gkac615Thoms, Buschauer, Ameismeier, Koepke, Denk et al., Structural basis for translational shutdown and immune evasion by the Nsp1 protein of SARS-CoV-2, Science,
doi:10.1126/science.abc8665Vora, Fontana, Mao, Leger, Zhang et al., Targeting stem-loop 1 of the SARS-CoV-2 5′ UTR to suppress viral translation and Nsp1 evasion, Proc Natl Acad Sci U S A,
doi:10.1073/pnas.2117198119Wang, Rimal, Tantray, Geng, Bhurtel et al., Prevention of ribosome collision-induced neuromuscular degeneration by SARS CoV-2-encoded Nsp1, Proc Natl Acad Sci U S A,
doi:10.1073/pnas.2202322119Ward, Bermingham, Ayoubkhani, Gethings, Pouwels et al., Risk of COVID-19 related deaths for SARS-CoV-2 Omicron (B.1.1.529) compared with Delta (B.1.617.2): retrospective cohort study, BMJ,
doi:10.1136/bmj-2022-070695Watson, Barnsley, Toor, Hogan, Winskill et al., Global impact of the first year of COVID-19 vaccination: a mathematical modelling study, Lancet Infect Dis,
doi:10.1016/S1473-3099(22)00320-6Xing, Ma, Tran, The DDX5/Dbp2 subfamily of DEAD-box RNA helicases, Wiley Interdiscip Rev RNA,
doi:10.1002/wrna.1519Xu, Cai, Ma, Jiang, Liu et al., The RNA helicase DDX5 promotes viral infection via regulating N6-methyladenosine levels on the DHX58 and NFκB transcripts to dampen antiviral innate immunity, PLoS Pathog,
doi:10.1371/journal.ppat.1009530Ye, Yuan, Chan, Zhang, Yu et al., Beneficial effect of combinational methylprednisolone and remdesivir in hamster model of SARS-CoV-2 infection, Emerg Microbes Infect,
doi:10.1038/s41592-022-01447-wYu, Van Berkel, Biessen, Therapeutic potential of VIVIT, a selective peptide inhibitor of nuclear factor of activated T cells, in cardiovascular disorders, Cardiovasc Drug Rev,
doi:10.1111/j.1527-3466.2007.00011.xYuan, Peng, Park, Hu, Devarkar et al., Nonstructural protein 1 of SARS-CoV-2 is a potent pathogenicity factor redirecting host protein synthesis machinery toward viral RNA, Mol Cell,
doi:10.1016/j.molcel.2020.10.034DOI record:
{
"DOI": "10.1128/mbio.00392-24",
"ISSN": [
"2150-7511"
],
"URL": "http://dx.doi.org/10.1128/mbio.00392-24",
"abstract": "<jats:title>ABSTRACT</jats:title>\n <jats:sec>\n <jats:title />\n <jats:p>SARS-CoV-2, the causative agent of COVID-19, has been intensely studied in search of effective antiviral treatments. The immunosuppressant cyclosporine A (CsA) has been suggested to be a pan-coronavirus inhibitor, yet its underlying mechanism remained largely unknown. Here, we found that non-structural protein 1 (Nsp1) of SARS-CoV-2 usurped CsA-suppressed nuclear factor of activated T cells (NFAT) signaling to drive the expression of cellular DEAD-box helicase 5 (DDX5), which facilitates viral replication. Nsp1 interacted with calcineurin A (CnA) to displace the regulatory protein regulator of calcineurin 3 (RCAN3) of CnA for NFAT activation. The influence of NFAT activation on SARS-CoV-2 replication was also validated by using the Nsp1-deficient mutant virus. Calcineurin inhibitors, such as CsA and VIVIT, inhibited SARS-CoV-2 replication and exhibited synergistic antiviral effects when used in combination with nirmatrelvir. Our study delineated the molecular mechanism of CsA-mediated inhibition of SARS-CoV-2 replication and the anti-SARS-CoV-2 action of calcineurin inhibitors.</jats:p>\n </jats:sec>\n <jats:sec>\n <jats:title>IMPORTANCE</jats:title>\n <jats:p>Cyclosporine A (CsA), commonly used to inhibit immune responses, is also known to have anti-SARS-CoV-2 activity, but its mode of action remains elusive. Here, we provide a model to explain how CsA antagonizes SARS-CoV-2 through three critical proteins: DDX5, NFAT1, and Nsp1. DDX5 is a cellular facilitator of SARS-CoV-2 replication, and NFAT1 controls the production of DDX5. Nsp1 is a viral protein absent from the mature viral particle and capable of activating the function of NFAT1 and DDX5. CsA and similar agents suppress Nsp1, NFAT1, and DDX5 to exert their anti-SARS-CoV-2 activity either alone or in combination with Paxlovid.</jats:p>\n </jats:sec>",
"alternative-id": [
"10.1128/mbio.00392-24"
],
"assertion": [
{
"group": {
"label": "Publication History",
"name": "publication_history"
},
"label": "Received",
"name": "received",
"order": 0,
"value": "2024-02-06"
},
{
"group": {
"label": "Publication History",
"name": "publication_history"
},
"label": "Accepted",
"name": "accepted",
"order": 1,
"value": "2024-02-08"
},
{
"group": {
"label": "Publication History",
"name": "publication_history"
},
"label": "Published",
"name": "published",
"order": 2,
"value": "2024-02-27"
}
],
"author": [
{
"affiliation": [
{
"name": "School of Biomedical Sciences, The University of Hong Kong, Pokfulam, Hong Kong"
}
],
"family": "Lui",
"given": "Wai-Yin",
"sequence": "first"
},
{
"affiliation": [
{
"name": "School of Biomedical Sciences, The University of Hong Kong, Pokfulam, Hong Kong"
}
],
"family": "Ong",
"given": "Chon Phin",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "School of Biomedical Sciences, The University of Hong Kong, Pokfulam, Hong Kong"
}
],
"family": "Cheung",
"given": "Pak-Hin Hinson",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "School of Biomedical Sciences, The University of Hong Kong, Pokfulam, Hong Kong"
}
],
"family": "Ye",
"given": "Zi-Wei",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "School of Biomedical Sciences, The University of Hong Kong, Pokfulam, Hong Kong"
}
],
"family": "Chan",
"given": "Chi-Ping",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-1921-5824",
"affiliation": [
{
"name": "Department of Microbiology, The University of Hong Kong, Pokfulam, Hong Kong"
}
],
"authenticated-orcid": true,
"family": "To",
"given": "Kelvin Kai-Wang",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "School of Biomedical Sciences, The University of Hong Kong, Pokfulam, Hong Kong"
},
{
"name": "School of Nursing, Tung Wah College, Kowloon, Hong Kong"
}
],
"family": "Yuen",
"given": "Kit-San",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-2778-3530",
"affiliation": [
{
"name": "School of Biomedical Sciences, The University of Hong Kong, Pokfulam, Hong Kong"
}
],
"authenticated-orcid": true,
"family": "Jin",
"given": "Dong-Yan",
"sequence": "additional"
}
],
"container-title": "mBio",
"container-title-short": "mBio",
"content-domain": {
"crossmark-restriction": true,
"domain": [
"journals.asm.org"
]
},
"created": {
"date-parts": [
[
2024,
2,
27
]
],
"date-time": "2024-02-27T14:01:11Z",
"timestamp": 1709042471000
},
"deposited": {
"date-parts": [
[
2024,
2,
27
]
],
"date-time": "2024-02-27T14:01:26Z",
"timestamp": 1709042486000
},
"editor": [
{
"affiliation": [],
"family": "Meng",
"given": "Xiang-Jin",
"sequence": "additional"
}
],
"funder": [
{
"award": [
"2021YFC0866100, 2023YFC3041600"
],
"name": "MOST | National Key Research and Development Program of China"
},
{
"DOI": "10.13039/501100002920",
"award": [
"C7142-20GF, T11-709/21-N"
],
"doi-asserted-by": "crossref",
"name": "Research Grants Council, University Grants Committee"
}
],
"indexed": {
"date-parts": [
[
2024,
2,
28
]
],
"date-time": "2024-02-28T00:42:09Z",
"timestamp": 1709080929673
},
"is-referenced-by-count": 0,
"issued": {
"date-parts": [
[
2024,
2,
27
]
]
},
"language": "en",
"license": [
{
"URL": "https://creativecommons.org/licenses/by/4.0/",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2024,
2,
27
]
],
"date-time": "2024-02-27T00:00:00Z",
"timestamp": 1708992000000
}
},
{
"URL": "https://journals.asm.org/non-commercial-tdm-license",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2024,
2,
27
]
],
"date-time": "2024-02-27T00:00:00Z",
"timestamp": 1708992000000
}
}
],
"link": [
{
"URL": "https://journals.asm.org/doi/pdf/10.1128/mbio.00392-24",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://journals.asm.org/doi/pdf/10.1128/mbio.00392-24",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "235",
"original-title": [],
"prefix": "10.1128",
"published": {
"date-parts": [
[
2024,
2,
27
]
]
},
"published-online": {
"date-parts": [
[
2024,
2,
27
]
]
},
"publisher": "American Society for Microbiology",
"reference": [
{
"DOI": "10.1016/S0140-6736(20)30154-9",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_2_2"
},
{
"DOI": "10.1038/s41591-022-01882-4",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_3_2"
},
{
"DOI": "10.1126/science.abl8506",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_4_2"
},
{
"DOI": "10.1016/S1473-3099(22)00320-6",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_5_2"
},
{
"DOI": "10.1136/bmj-2022-070695",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_6_2"
},
{
"DOI": "10.1002/jmv.26140",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_7_2"
},
{
"DOI": "10.1016/j.celrep.2022.110954",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_8_2"
},
{
"DOI": "10.1073/pnas.2117198119",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_9_2"
},
{
"DOI": "10.1093/nar/gkad269",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_10_2"
},
{
"DOI": "10.1093/nar/gkac615",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_11_2"
},
{
"DOI": "10.1101/gad.350829.123",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_12_2"
},
{
"DOI": "10.1016/j.molcel.2023.09.002",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_13_2"
},
{
"DOI": "10.1073/pnas.2202322119",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_14_2"
},
{
"DOI": "10.1073/pnas.2017715118",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_15_2"
},
{
"DOI": "10.1080/15476286.2021.1999103",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_16_2"
},
{
"DOI": "10.1016/j.chom.2021.01.015",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_17_2"
},
{
"DOI": "10.1038/s41594-020-0511-8",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_18_2"
},
{
"DOI": "10.1126/science.abc8665",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_19_2"
},
{
"DOI": "10.1038/s41467-022-34571-4",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_20_2"
},
{
"DOI": "10.1016/j.celrep.2021.109841",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_21_2"
},
{
"DOI": "10.1128/jvi.01865-22",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_22_2"
},
{
"DOI": "10.1371/journal.ppat.1002331",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_23_2"
},
{
"DOI": "10.53730/ijhs.v6nS6.10793",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_24_2"
},
{
"DOI": "10.3389/fimmu.2020.00195",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_25_2"
},
{
"DOI": "10.1186/1478-811X-7-25",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_26_2"
},
{
"DOI": "10.1002/btm2.10436",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_27_2"
},
{
"DOI": "10.1128/spectrum.01504-21",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_28_2"
},
{
"DOI": "10.1016/j.eclinm.2020.100591",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_29_2"
},
{
"DOI": "10.1016/j.bcp.2019.03.022",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_30_2"
},
{
"DOI": "10.1128/JVI.01352-15",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_31_2"
},
{
"DOI": "10.1128/JVI.00266-21",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_32_2"
},
{
"DOI": "10.1002/jmv.28057",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_33_2"
},
{
"DOI": "10.1016/j.molcel.2020.10.034",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_34_2"
},
{
"DOI": "10.1002/wrna.1519",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_35_2"
},
{
"DOI": "10.1128/jvi.00002-22",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_36_2"
},
{
"DOI": "10.1371/journal.ppat.1009530",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_37_2"
},
{
"DOI": "10.1111/j.1527-3466.2007.00011.x",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_38_2"
},
{
"DOI": "10.1016/j.tcb.2010.09.011",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_39_2"
},
{
"DOI": "10.7554/eLife.74877",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_40_2"
},
{
"DOI": "10.1097/01.qco.0000172698.52408.be",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_41_2"
},
{
"DOI": "10.1053/j.ajkd.2022.01.001",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_42_2"
},
{
"DOI": "10.1164/rccm.202108-1830LE",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_43_2"
},
{
"DOI": "10.1016/j.celrep.2021.108959",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_44_2"
},
{
"DOI": "10.1021/acschemneuro.1c00019",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_45_2"
},
{
"DOI": "10.1038/s41598-019-46592-z",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_46_2"
},
{
"DOI": "10.1161/HYPERTENSIONAHA.116.07140",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_47_2"
},
{
"DOI": "10.1371/journal.ppat.1009225",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_48_2"
},
{
"DOI": "10.1167/iovs.04-1022",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_49_2"
},
{
"DOI": "10.1016/j.cmi.2023.01.002",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_50_2"
},
{
"DOI": "10.1111/j.1442-9071.2008.01784.x",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_51_2"
},
{
"DOI": "10.1002/iid3.509",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_52_2"
},
{
"DOI": "10.1038/s41598-022-14194-x",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_53_2"
},
{
"DOI": "10.3390/ph15091039",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_54_2"
},
{
"DOI": "10.3389/fcell.2021.667879",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_55_2"
},
{
"DOI": "10.1080/22221751.2021.1885998",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_56_2"
},
{
"DOI": "10.1038/s41592-022-01447-w",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_57_2"
},
{
"DOI": "10.1182/blood-2009-07-228387",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_58_2"
},
{
"DOI": "10.1002/1521-4141(2002010)32:10<2971::AID-IMMU2971>3.0.CO;2-G",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_59_2"
},
{
"DOI": "10.1038/s41423-022-00855-4",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_60_2"
},
{
"DOI": "10.1371/journal.ppat.1011186",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_61_2"
},
{
"DOI": "10.1093/cid/ciac203",
"doi-asserted-by": "publisher",
"key": "e_1_3_4_62_2"
}
],
"reference-count": 61,
"references-count": 61,
"relation": {},
"resource": {
"primary": {
"URL": "https://journals.asm.org/doi/10.1128/mbio.00392-24"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [
"Virology",
"Microbiology"
],
"subtitle": [],
"title": "Nsp1 facilitates SARS-CoV-2 replication through calcineurin-NFAT signaling",
"type": "journal-article",
"update-policy": "http://dx.doi.org/10.1128/asmj-crossmark-policy-page"
}